We wrote this post as a guest post for the rOpenSci blog, where it was originally reviewed and published.
Imagine you are a fish ecologist who compiled a list of fish species for your country. 🐟
Your list could be useful to others, so you publish it as a supplementary file to an article or in a research repository. That is fantastic, but it might be difficult for others to discover your list or combine it with other lists of species. Luckily there’s a better way to publish species lists: as a standardized checklist that can be harvested and processed by the Global Biodiversity Information Facility (GBIF). We created a documented template to do that in R, which recently won the GBIF Ebbe Nielsen Challenge. In this post we explain how we did that and highlight some of the tools we discovered along the way.
What is GBIF?
For those unfamiliar with the Global Biodiversity Information Facility (GBIF), it is an international network and research infrastructure funded by the world’s governments, aimed at providing anyone, anywhere, open access to data about all types of life on Earth. GBIF is best known for the trove of species occurrences (over 1 billion!) it is making accessible from hundreds of publishing institutions, but it is doing the same for species information, such as names, classifications and known distributions.
Anyone can publish species information (called “checklist data”) to GBIF. When you do, GBIF will create a page for your dataset (example), assign a DOI, and match your scientific names to its backbone taxonomy, allowing your data to be linked, discoverable and integrated. Species pages on GBIF for example (example) are automatically built from the over 25,000 checklist datasets that have been published. All your checklist information also becomes available via the GBIF API (example) and can be queried using the rgbif package.
So, why isn’t everyone publishing checklists? Because the data can only be reasonably integrated if they are published in a standardized way. All GBIF mediated-data (including occurrences) have to be published in the Darwin Core standard, fitting a standard structure (Darwin Core Archives), columns (Darwin Core terms) and sometimes values (controlled vocabularies), which can be challenging. Templates and the GBIF Integrated Publishing Toolkit facilitates standardization or “mapping”, but only caters for the most basic use cases. It also forces you to change the structure of your source data and involves many manual steps. To solve this, we created a recipe to facilitate and automate this process using R.
The checklist recipe
Our Checklist recipe is a template GitHub repository for standardizing species checklist data to Darwin Core using R. It contains all the ingredients to make your data standardization open, repeatable, customizable and documented. The recipe has considerably streamlined our own work to publish seven checklists on alien species for Belgium, which is one of the goals of the Tracking Invasive Alien Species (TrIAS) project, an open data-driven framework to support Belgian federal policy on invasive species. Making biodiversity research more efficient and reproducible is the core mission of our team at the Research Institute for Nature and Forest (INBO). A mission we tackle by supporting researchers in publishing open data and developing open source software.
The basic idea behind the Checklist recipe is:
source data → Darwin Core mapping script → generated Darwin Core files
By changing the source data and/or the mapping script, you can alter the generated Darwin Core files. The main advantage is repeatability: once you have done the mapping, you don’t have to start from scratch if your source data has been updated. You can just run the mapping script again (with a little tweak here and there) and upload the generated files to a GBIF Integrated Publishing Toolkit (IPT) for publication. And by having a mapping script, your mapping is also documented.
Rather than explaining how you can use the Checklist recipe - we’ve documented this in a wiki - we’d like to highlight some of the tools and techniques we discovered in developing it.
Cookiecutter data science
The recipe shares the same repository structure we use for all our data transformation repositories. We didn’t invent one, but adopted Cookiecutter Data Science: “a logical, reasonably standardized, but flexible project structure for doing and sharing data science work”. The main advantage we think is that it allows anyone (and us) to easily find their way around a repository, making it easier to contribute. It also saves precious time setting up a repository, because there are fewer decisions (e.g. naming things) to be made.
Below is the directory structure we adopted for checklist repositories. Files and directories indicated with GENERATED should not be edited manually.
├── README.md : Description of the repository
├── LICENSE : Repository license
├── checklist-recipe.Rproj : RStudio project file
├── .gitignore : Files and directories to be ignored by git
│
├── data
│ ├── raw : Source data, input for mapping script
│ └── processed : Darwin Core output of mapping script GENERATED
│
├── docs : Repository website GENERATED
│
└── src
├── dwc_mapping.Rmd : Darwin Core mapping script, core functionality of the repository
├── _site.yml : Settings to build website in /docs
└── index.Rmd : Template for website homepage
R Markdown
The core functionality of our recipe is an R Markdown file called dwc_mapping.Rmd (i.e. the “mapping script”). If you are unfamiliar with R Markdown, it is a file format that allows you to mix text (written in Markdown) with executable code chunks (in R). It is comparable with an R script, in which the comments explaining the code are given as much value as the code itself. It has the advantage that you can describe and then execute each step of your data processing in the same document, nudging you to better document what you are doing. This is called “literate programming” and it is one of the steps to make research more reproducible.
You can simply run the code of an R Markdown file by opening it in RStudio and choosing Run > Run all (or code chunk by code chunk) or you can render as a report using knit. R Markdown supports a whole range of file formats for these reports (including html and pdf).
R Markdown websites
If you are using R Markdown in a GitHub repository, you have all the ingredients to generate a small website showcasing your mapping script in a visually pleasing way (example). And it can be hosted on GitHub for free! To learn more, read the documentation on R Markdown websites. The basic setup is:
- Create an
index.Rmd file at the same level as your other R Markdown files (in the src directory). That file will be the homepage of your website. Since we don’t want to repeat ourselves, we inject the content of the repository README.md in the homepage.
- Create a
_site.yml file at the same level as your index.Rmd file. It contains the settings for your website. Set at minimum a name, navbar and output_dir: "../docs" so the website is created in the /docs directory (which you need to create as well).
- Go to
Build > Configure Build Tools… in RStudio and set Project build tools as Website with Site directory as src. You will now have a build pane in RStudio where you can click Build Website to build your website.
This setup has already been done in our recipe.
To serve your website, commit and push your changes to GitHub, go to your repository settings and choose the /docs directory to build a GitHub pages site. After a couple of seconds, your website should be available at <username>.github.io/<reponame>/. Don’t forget to add it to your repo description so users can find it.
here
In order to share working directory and build settings, we like to include the RStudio project file in our repositories, ideally in the root and with the same name as the project/repository (e.g. checklist-recipe.Rproj). But that posed a problem with relative links and the difference between running and knitting the code.
- When running code, the working directory is where the
.Rproj file is located (the root), so a relative path to a data file would be data/raw/checklist.xlsx
- When knitting/building code, the working directory is where the R Markdown file is located (
/src), so a relative path to a data file would be ../data/raw/checklist.xlsx
Obviously that created problems and the only way we could make it work is by having the .Rproj file in the /src directory, so that both running and knitting would use the same working directory. That is, until we discovered the here package:
Rather than hardcoding a path, you just use:
input_data <- read_excel(path = here("data", "raw", "checklist.xlsx"))
And here() will walk up your directory structure until it finds something that looks like the top-level and find the file from there. Makes linking to files so much easier!
The %<>% pipe
We use “piping” (i.e. using the pipe operator %>% or pipe) to increase the readability of our code:
# Take the dataframe "taxon", group the values of the column "input_kingdom" and show a count for each unique value
taxon %>%
group_by(input_kingdom) %>%
count()
Which is a more readable way than the classic approach of nesting functions:
count(group_by(taxon, input_kingdom))
One pipe you might not recognize is the compound assignment pipe operator or %<>%:
# Take the dataframe "taxon" and add the column "kingdom" with value "Animalia" for all records
taxon %<>% mutate(kingdom = "Animalia")
Which is a shorthand for writing:
taxon <- taxon %>% mutate(kingdom = "Animalia")
It’s mainly useful if you want to transform a dataframe in consecutive steps, like adding Darwin Core terms as columns. The %<>% pipe is not included with dplyr or tidyverse, so you have to load magrittr separately to use it:
rgbif
We mostly rely on the dplyr functions mutate(), recode() and case_when() to map data to Darwin Core. But to verify that our scientific names are well-formed, we use the rOpenSci package rgbif to interact with another service by GBIF: the name parser:
parsed_names <- input_data %>%
distinct(input_scientific_name) %>% # Retrieve unique scientific names
pull() %>% # Create vector from dataframe
rgbif::parsenames() # Parse scientific names and save as parsed_names
The name parser checks if a scientific name (a string such as Amaranthus macrocarpus Benth. var. pallidus Benth.) is well-formed (i.e. follows the nomenclatural rules) and breaks it down in components:
- type:
SCIENTIFIC
- genusOrAbove:
Amaranthus
- specificEpithet:
macrocarpus
- infraspecificEpithet:
pallidus
- authorship:
Benth.
- canonicalName:
Amaranthus macrocarpus pallidus
- rankMarker:
var.
We use this information to verify if our scientific names are indeed written as scientific names and to populate the taxon rank (a mandatory Darwin Core term for checklists) using rankMarker. Note that the name parser does not check the existence of a scientific name against an existing registry. That is done by the GBIF species lookup tool we discussed above, which verifies the existence of a name in the GBIF backbone taxonomy.
rgbif provides many more functions to interact with the Global Biodiversity Information Facility and we use the package extensively for our TrIAS project.
Conclusion
Our recipe grew organically from experience we gained publishing data to GBIF. We saw the GBIF Ebbe Nielsen Challenge as an opportunity to bottle and document what we had learned in an opinionated template to help others and we hope this blog post highlighted a few tips and tricks that might be useful to you as well. If you want to use the recipe to publish your own checklist data, start here.
We are strongly convinced that the future of biodiversity research (and science in general) is open. We are proud to co-win the GBIF Ebbe Nielsen Challenge and took it as an opportunity to give back. That is why we are donating half of our prize money to NumFOCUS, an organization sponsoring several open source initiatives we rely on every day (including rOpenSci) improving the quality of science worldwide. Supporting open source research software means supporting your own research after all.
If you want to get in touch with our team, contact us via Twitter or email.
]]>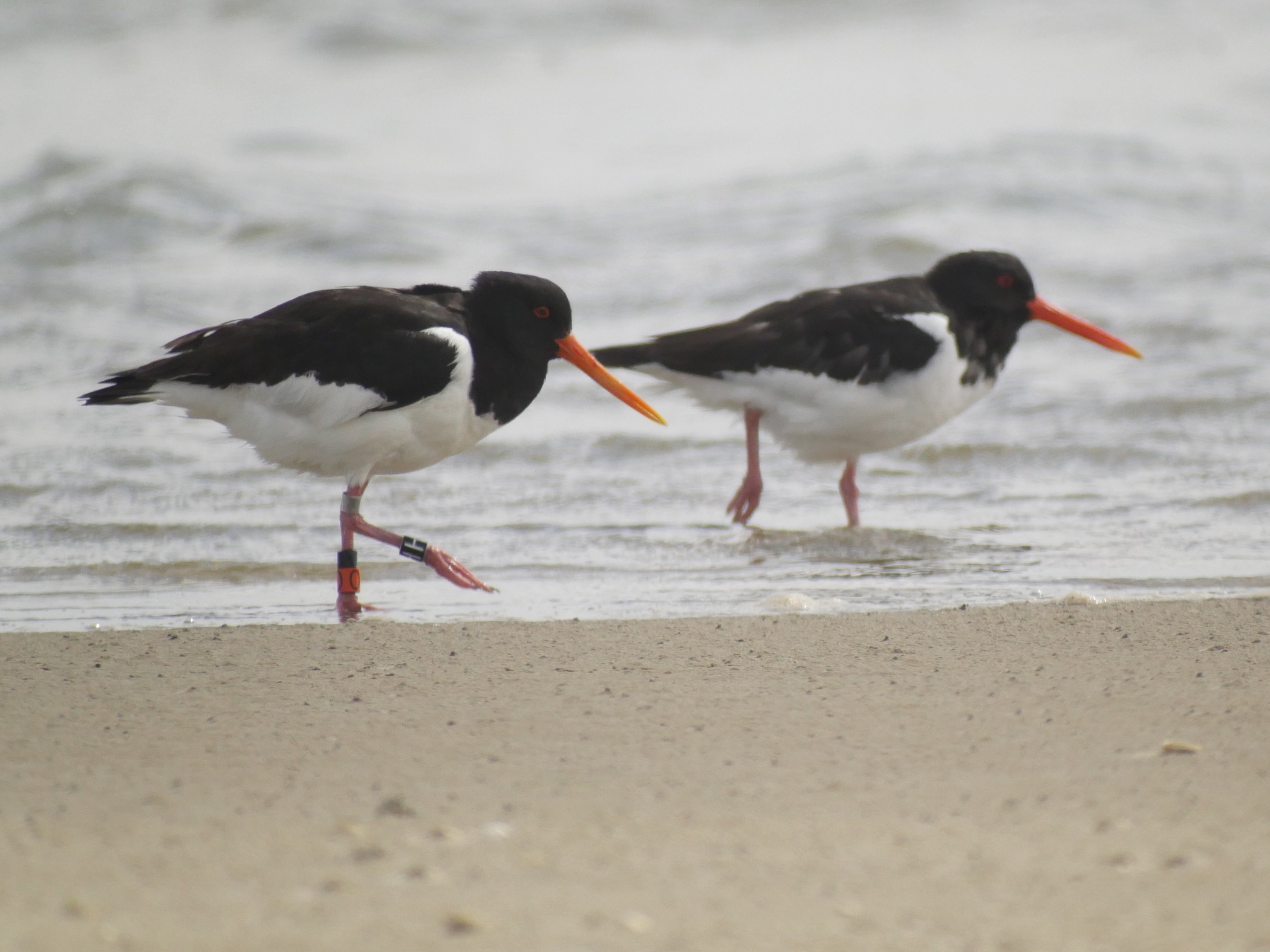 Oystercatchers on Vlieland, the Netherlands. The left individual is equipped with a GPS tracker and colour rings.
Oystercatchers on Vlieland, the Netherlands. The left individual is equipped with a GPS tracker and colour rings.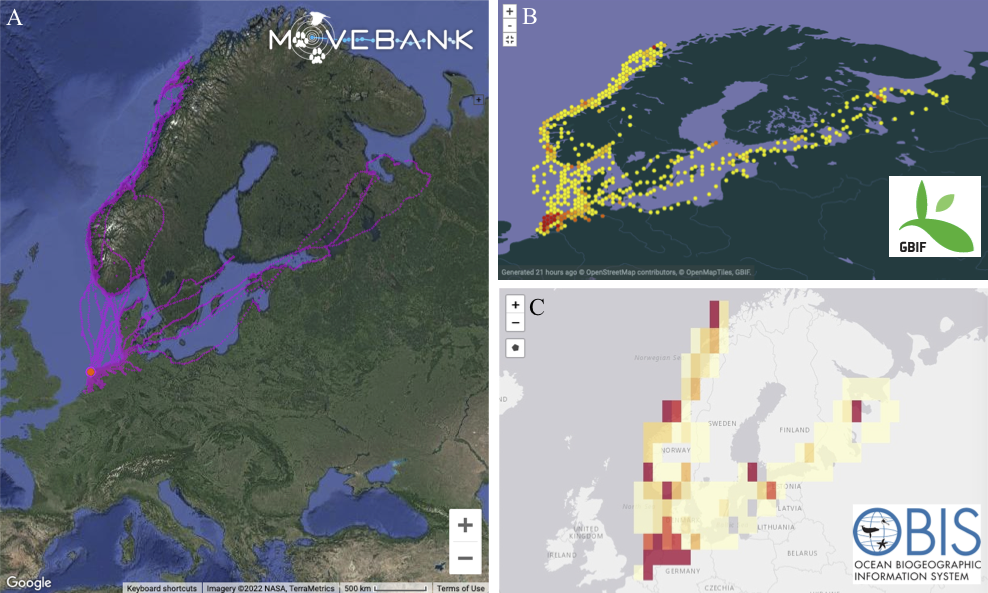 Oystercatcher GPS tracks from Vlieland are available via Movebank, GBIF and OBIS.
Oystercatcher GPS tracks from Vlieland are available via Movebank, GBIF and OBIS.